CSReport : Search, identification and quantification of CSR junctions in high-throughput sequencing data
Contact
Sophie Péron, CR INSERM
J Immunol. 2017 May 15;198(10):4148-4155.
CSReport: A New Computational Tool Designed for Automatic Analysis of Class Switch Recombination Junctions Sequenced by High-Throughput Sequencing.
Boyer F, Boutouil H, Dalloul I, Dalloul Z, Cook-Moreau J, Aldigier JC, Carrion C, Herve B, Scaon E, Cogné M, Péron S.
The CSReport notebook file (ZIP archive) including the reference files is available on request by email to Sophie Péron.
Class switch recombination…
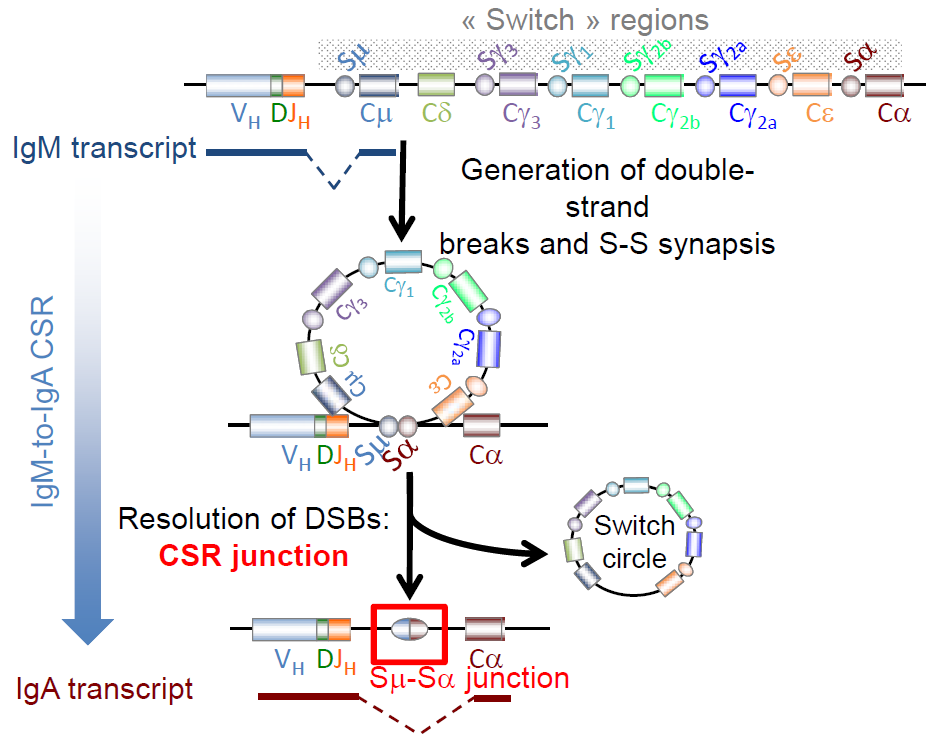
During their terminal maturation, B cells can be induced to class-switch recombination (CSR) leading to immunoglobulin heavy chain (IgH) locus rearrangement :
- – double-strand breaks (DSBs) are generated in switch regions,
- – joining and repair of DSBs leads to the expression of a different isotype.
Elucidation of CSR ‘‘switch-switch’’ junctions allows to trace back recombination events and to infer break/repair mechanisms and pathways.
Specific amplification and sequencing are the key techniques to get break points’ positions and junctions’ structures. The classical method relies on Sanger sequencing and fastidious manual ‘‘BLAST-like’’ analysis.
… with a deeper insight
High-throughput sequencing (HTS) will allow a better characterization of CSR in large cell population and open the way for quantification.
Using HTS-based experimental protocol, we generate millions of sequencing reads per sample that may support a class switch recombination event.
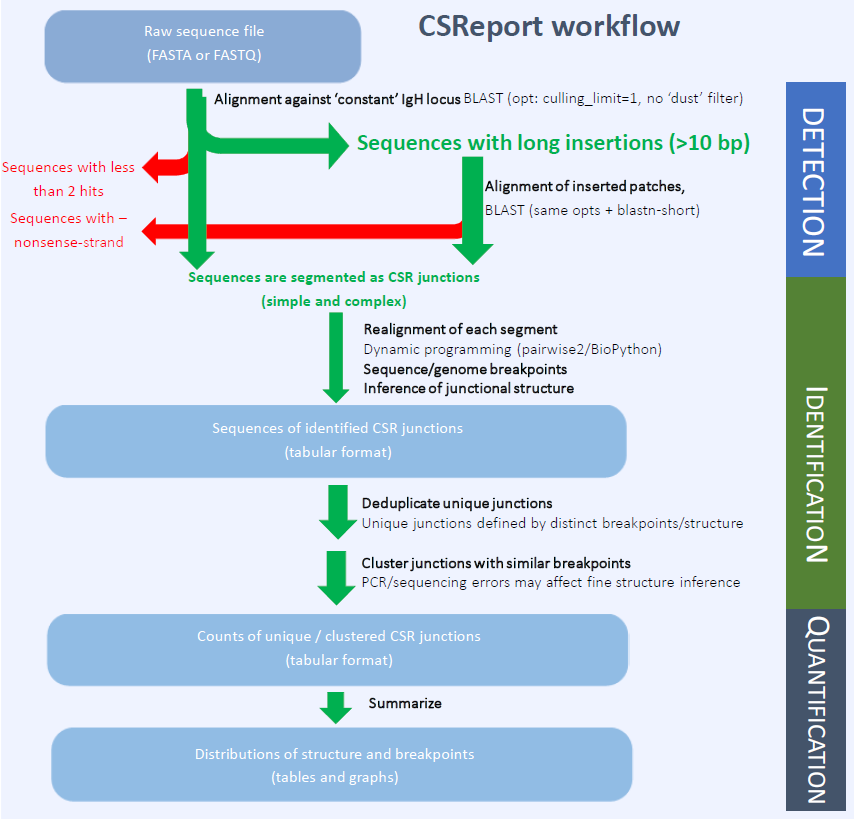
To achieve optimal use from these large datasets, we developed CSReport, a new computational tool which automatically identifies and summarizes sequences that support recombination between two switch regions of the IgH locus.
Results
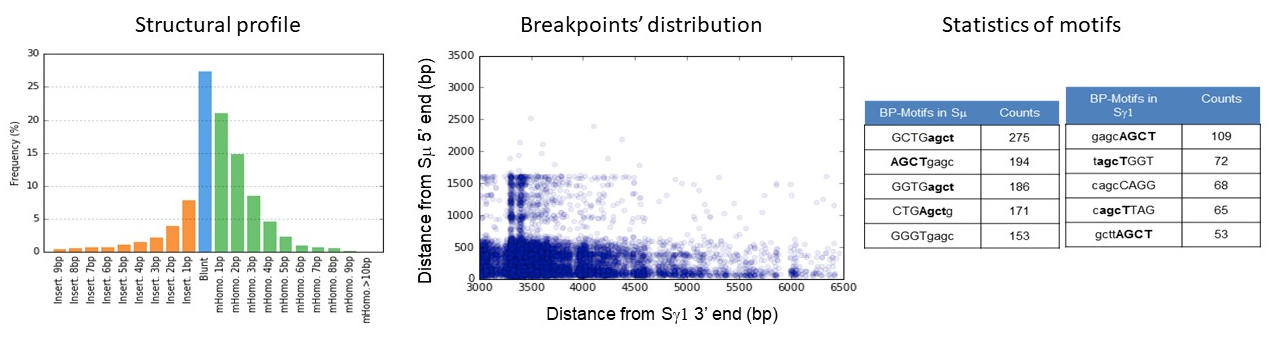
CSReport allows identification of junction from two pairwise alignments. Break points are determined with 1-bp resolution for accurate structure assignment. CSReport describes the structural profile of the junctions (Identification of junction structure with insertion, with homology or blunt junctions), the distribution of junction breakpoints in the sequence references and the most frequent 8nt-motifs at breakpoints in switch sequences.
CSReport requires Python3 and Jupyter environments (preferably from an Anaconda distribution) with updated Biopython package. Raw sequencing data should be single-end reads (in multi-fasta or fastq formats). CSReport works with a custom reference (derived from NCBI sources) of a constant IgH locus sequence and annotations (humans or mice). The reference should be chosen according to the model organism. Running time (laptop) ≈ 30 mins/106 sequences.